P.C. Schreckenberger's Microbiology Homepage
.Introducing the ASHEX Encyclopedia and ASHEX24XB WIP by A.P. Schreckenberger – April 27, 2024
Introducing the 1st Edition of The ASHEX Encyclopedia. This document cements the project's policy towards transparency by publishing 35.26% of the raw biochemical data sheets that forged the ASHEX identification matrix. These sheets were provided by Loyola University Medical Center after being scrubbed of personal information, and thanks to Dr. Amanda Harrington, Kathleen McKinley, and all the staff at LUMC for facilitating my dream to unleash a true Doctors Schreckenberger publication.
In addition to containing summary tables for the entire ASHEX23 matrix, every species represented by the table, and the raw scans of over 350 biochemical sheets, The ASHEX Encyclopedia provides a full mathematical description of how the Web Identification Programs (WIPs) work, an overview of the data sources involved, a summary of interpreting WIP results, our transparency statement and policy, and my casual commentary.
Barring the rediscovery of other biochemical sheets used to populate the matrix, The ASHEX Encyclopedia truly is the ultimate deliverable of the SCHNF ASHEX Project, and I personally view it as the culmination of our efforts. PDF downloads are available on the Non-Fermenter ID page in the Matrix & Encyclopedia section. Please note that the files are quite large. The uncompressed file is almost 1 GB, and the compressed file is around 200 MB.
Furthermore, I have introduced an update to the ASHEX WIP. The ASHEX24XB program still utilizes Version 23 of the matrix. However, while compiling the encyclopedia, I learned more about my dad's research process, and it made me yearn for a new feature. This new feature has now been introduced. At the bottom of the ASHEX form, users can specify a genus to display along with the standard calculator output. Perhaps your intuition is pointing you in a certain direction, or maybe you want to keep tabs on modal scores as you switch on the Wilson Binomial table. I believe this tool will be beneficial -- without adding too much complication.

GENERAL ARCHIVE
April 2024: The ASHEX Encyclopedia
July 2023: ASHEX23XB Deployment
September 2022: The New Site
February 2022: schreckenbergeri
January 2022: Vibrio Bug Fix!
February 2020: Dr. Elmer Koneman
July 2019: Update Question
December 2017: Night to Remember
September 2017: ASHEX18XB WIP!
September 2017: Tutorial Video
December 2016: ASHEX16X Inbound
December 2016: Dr. Paul's Passing
March 2014: ASHEX14X
February 2014: Code Rewrite
November 2012: The Magi
July 2012: The WIP is born!
May 2011: Welcome to PSCHRECK
.ASHEX23XB Deployment + Major Backend Updates by A.P. Schreckenberger – July 27, 2023
Five years have passed since I last touched the calculator code, but significant updates have impacted the language used to drive the tools. As I mentioned when I built the new site, carrying around things that can't be updated to work with modern software releases are security liabilities, and this mindset extends to all of the WIPs. I will be rolling out major upgrades 'under the rug' in the near future. While I don't expect there to be any impacts to users, there have been noteworthy shifts in how certain variables were processed in the existing code, so please reach out if you see any errors or warnings that arise during use. I have tested things in a build on my machine, so I expect all the WIPs to run smoothly, but one tester is never enough. Please keep in mind that this update will affect all of the WIPs on the site, including the ASHEX, Enterics, Vibrio, MAGI, and Salmonella WIPs. You may need to clear your browser cache if you are still seeing links to ASHEX18.
I have also been sitting on a bit of a surprise. The ASHEX23X matrix and its associated WIP will be available after the rollout. ASHEX23 represents 109 taxa with biochemical results from 1001 clinical isolates. Since the WIPs were launched in 2012, over 28000 identifications have been made with this free-to-use, open-data system, and countless more identifications were made with the previous tables in the eight years before the first WIP.
As always, the Microbiology community means a lot to me given my dad's history, and I have always regarded this personal project of ours as my most important scientific contribution.
May life treat you all well!
~Adam
.The New PSCHRECK.com – September 14, 2022
Hello, everyone! Welcome to the new PSCHRECK.com webpage. First thing first, if you are looking for the links to the WIPs and you are on a narrow/mobile window screen, said WIP links are located in the sidemenu. You can access the menu by clicking the button in the upper right corner of the page (indicated by the red arrow in the picture below). That action will display the familiar links on the left side of your screen.
With that out of the way, now I can answer why I moved away from the previous site. The third iteration of PSCHRECK.com, which launched in May 2011, ran on WordPress. This was a major step forward over the 2nd Generation Flash page and the original HTML one. WordPress also offered some great advantages when it came to my dad's intentions. He wanted to make regular blogs, and WordPress automatically link generates for new posts. He wanted an easy archive, and WordPress was great for that. The sites also tend to just look clean! However, in the last 11 years, HTML has come a long way. We now have CSS, which facilitates amazing style options, and in my opinion, WordPress leaves a lot to be desired these days. Dedicated WordPress hosting plans cost a pretty penny. The sites tend to be slow because you're without caching if the host does not subscribe to a caching plan. Finally, the software updates can be prohibitive. Every time a major WP update got pushed, I would fear updating the server. I have had to scramble to rebuild the site on multiple occasions, and the latest event, which led to me finally pulling the trigger on a relaunch, rendered me unable to reliably edit/post or view images on the site. When you hesitate to implement security updates on a piece of software because you're afraid your page will break, it's time to move on from that software.
The end result of the PSCHRECK Gen4 Initiative is a site that looks just as clean as the previous version, loads over 10X faster (yes, I measured the load times), no longer requires annoying software updates to maintain functionality and security, and--most importantly--continues to provide quality identification tools to the micro community. Over the years, particularly in the ones following my dad's death, I have reaffirmed my dedication to WIP availability and engagement. We're still here.
May your day be awesome,
Dr. A.P. Schreckenberger
.Franklinella schreckenbergeri – February 28, 2022
Congratulations to Kathryn A. Bernard, Ana Luisa Pacheco, Tamara Burdz, Deborah Wiebe, and Anne-Marie Bernier for their recently-published article in the International Journal of Systematic and Evolutionary Microbiology [DOI 10.1099/ijsem.0.005247]. In the manuscript, they describe the assignment of provisionally named CDC group NO-1 strains. One of those strains was named Franklinella schreckenbergeri in honor of my father and in memory of his contributions to the field of Microbiology.
Such an honor is definitely appreciated by my family. Thank you for keeping his legacy alive.
.Vibrio WIP Bug Fix! – January 28, 2022
Melanie Orth at MDH pointed out a peculiarity in the Vibrio WIP. Two rows were labelled as Grimontia hollisae, and this was throwing a wrench in a Vibrio metschnikovii identification. The labelling error is one that existed even before I built the table. It is actually in the source documentation that I received years ago, so this one is definitely a gold-star bug find.
Using the Koneman Atlas, I verified that one of the duplicated rows does, in fact, match Vibrio metschnikovii, and I updated the species list to reflect this. Melanie reran the lab isolate, and it came back 100% Vibrio metschnikovii as expected.
Again, thanks for bringing this one to my attention. It’s nice to put on the micro hat for a day.
.Passing of Dr. Elmer Koneman – February 23, 2020
I would like to offer my condolences to the family and friends of Dr. Elmer Koneman, who passed away on the 14th of February. Many people in the field will remember the amazing contributions Elmer gave, his presence, and his guidance. My dad revered Elmer, and it would be impossible for me to count the ways his life was impacted by Elmer’s energy.
My life was also touched by Elmer. He always welcomed me to the lab when he was still at UIC. My first presentation as an aspiring scientist-to-be was in the 90s (I want to say 96) at Micro in the Mountains. Breckenridge is etched upon the memories of my childhood. And, I have to mention that I have always had a copy of THE BOOK because so many people I admired put it together.
This is a monumental loss, and yet, we have to keep going. Stand on their shoulders and keep on shining.
.A Serious Question... – July 17, 2019
Been a while everyone! Happy 2019! Looking over the ASHEX usage data, I’ve spotted something interesting that I’d like to bring up. It seems that a significant percentage of users are turning to the ASHEX14 matrix for identification. On its face, I can understand why people are doing this. The legacy version of the ID tool was the last one that I worked on with my father before his passing.
That being said, there are some substantial reasons to use the ASHEX18 matrix instead. When my father passed away, he was wrapping up a project to check the identifications in the ASHEX table against MALDI-TOF and gene sequencing. At the time, there were 900+ isolates in the dataset, and from those, 21 “problem children” were found. These 21 cases showed discrepancies between the three methods, and all of them were resolved thanks to the careful work from the lab. The ASHEX18 version has these improvements along with the Wilson Binomial Tool.
So my serious question comes from trying to understand the persistence in legacy version use. I genuinely believe that the improvements in ASHEX18 really warrant the use of the current matrix. Anyhoo! Hope everyone is doing alright. Feel free to contact me if you feel so inclined. I’m always happy to chat.
~Adam
.Congrats Jeanne – December 8, 2017
Last night, I had the honor of attending the retirement celebration of Jeanne vonRentzell, an undoubtedly boisterous voice in the Micro world at Loyola. I didn’t talk about it last night because the event was already marked by the joyous train of speeches, but I do have a story to share. My dad and I would often talk about things going on in our labs. People were no exception, and there were always good tales regarding growth and the future — and how good a place the lab happened to be in.
One of the moments that stands out to me when it comes to Jeanne was the trip to Boston. This was an incredibly hectic time in the Schreckenberger House. The week before, we had just gone to Minnesota so I could get a place to live. I bought a car, and I was officially moving in two weeks. Of course, in the middle of that two week span was the ASM meeting, and my dad wanted me to present ASHEX in his GNNFB session. That certainly brought about its own chain of pride, but the one thing that we was undeniably stoked about was Jeanne’s poster.
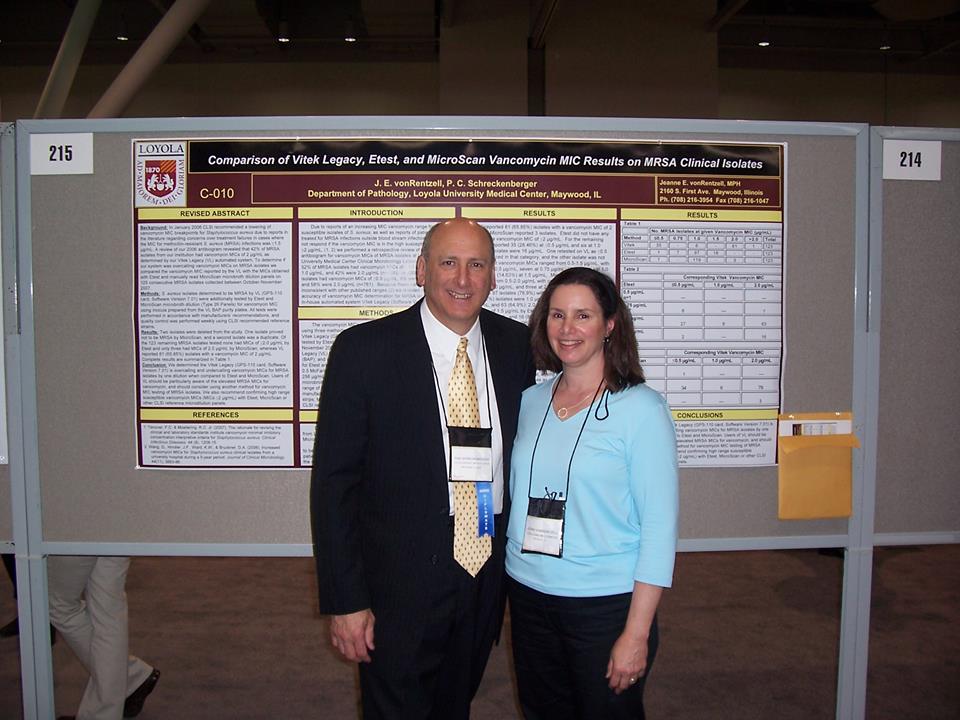

Oh my gosh, he was unfathomably excited about this poster. He ate it up and was beyond happy when it came to the Masters push. That’s just the kind of guy he was, and how could you not sit there and eat it up? He was proud of everyone who worked with him, and that snapshot barely captures a fraction of how proud he was when you, Jeanne, presented that work.
If you thought I couldn’t get sappier… the lab had a surprise for me last night as well. When the 1000th clinical isolate was entered into ASHEX, I knew there had to be a party. That lab is filled with party people. You don’t just hit a milestone. It worked out that the retirement party was at Giordano’s and that was also my first choice for a little get-together. Things converged. Celebrate!

I guess I shouldn’t have been surprised at all that Roman and Kathy called me up and presented me with the certificate above. There isn’t a whole lot I can do to express the feels on that one. The WIPs are probably the best things I have created, even with all the physics stuff factored in. This system has helped people (a heck of a lot of people), and it would not have been possible to build without the work of the lab. It’s a great team accomplishment, and it’s not going anywhere as long as I’m around to maintain it.
~Adam
.ASHEX18XB WIP is now live! – September 17, 2017
Greetings Microbiology Community,
I come to you today with a special announcement. Thirteen years after my dad first approached me with the idea of building the ASHEX table, the 1000th Gram-negative, non-fermenting clinical isolate was recorded this past week. The additional biochemical results are stored in the brand new ASHEX18XB Web Identification Program, which has now replaced the link in the menu at the top of the page.
The corresponding ASHEX18X matrix will be available for download shortly in the Non-Fermenter ID section of the site, and it represents everything my dad put into this field and more. When my father passed away in November, we had 932 isolates recorded in the table. After visiting his office, I found an additional 39 isolates with identifications he had checked and approved, which brought the total isolate count to 971 in the 17X version.
There were additional mysteries uncovered. Two of his colleagues, Kathy McKinley and Joyce Tjhio had performed numerous tests on these organisms over the years, and a comprehensive comparison of biochemical, MALDI-TOF, and gene sequencing techniques had been carried out in the months prior to that November. But the stars seemed to align, and the rollout of 17X went on without a hitch.
After speaking with my dad’s successor, Dr. Amanda Harrington, she reaffirmed an effort that had been a fantasy for me. We’d get the table to 1000 clinical isolates, and now we have.
It is hard for me to put into words my feelings on this one. He should have been here for this, and he is not. But in a way, he has been here the whole time. When we were worried a 13-year-old sheet had been misplaced, Kathy would somehow find it on top of a stack in the lab. If I grew curious about a result, the answer seemed to be literally written out in his chapter of The Koneman. Every step carried a little bit of magic, and that bridge that this project forged between the worlds of physics and microbiology grew stronger with each question and answer.
It is through the strength and determination of the colleagues that loved my dad and shared in his joyous life that I can say the following to you now:
Today, ASHEX18XB is now live! The matrix represents 108 taxa with biochemical results from 1000 clinical isolates. Since the WIPs were launched in 2012, over 18500 identifications have been made with this free-to-use, open data system, and countless more identifications were made with the table in the eight years before that.
A special thank you to those at UIC, Loyola, and elsewhere that touched Dr. S’s life.
Respectfully,
Dr. Adam P. Schreckenberger, Ph.D.
.ASHEX Tutorial Video – September 12, 2017
Greetings everyone, Adam here with an end-of-summer announcement. An updated version of the ASHEX identification matrix is in the works. The folks at Loyola University Medical Center are helping me complete the push to 1000 clinical isolates, a mark that my dad and I always viewed as the pipe dream benchmark. During this work, I realized that I haven’t made a tutorial video since 2014, and the more I thought about it, the more I convinced myself that it was time for an update.
.ASHEX16X, Life Carries On – December 9, 2016
ASHEX16X is now live! This update was built using a stack of biochemical test results my dad (Dr. Paul) had left for me to enter before his passing. Some of you might have also seen that there are now two ASHEX (GNNFB) links. I have decided to freeze ASHEX14X and keep it available. While I — and the great group at Loyola — are 110% committed to updating the ASHEX table to the highest caliber of standards, the caliber to which my father demanded, I realize there are some out there who might not be comfortable using tools built in the wake of his expertise. The WIP as it was at the time of his passing (ASHEX14X) will remain available, while the other link (ASHEX16X) will carry future developments.
Right now, there are several updates in the works. I improved the appearance of the Enterics WIP form last night. I plan to do the same to the Vibrio WIP, as well. Like a good teacher, my dad left behind NFB sheets that pose interesting questions, and I aim to get that sorted with the help of his lab. In other words, the update to ASHEX16X is the first step in a two-step upgrade. ASHEX17X will follow shortly (likely early 2017). This is the reason why the raw 16X table file is not currently available for download like the 14X one is. I’m moving forward immediately. It is a Schreckenberger trait.
Also, the typical note: ASHEX16X represents 104 taxa using data from 932 clinical isolates. Thanks for your energy and support!
~Adam
.The Passing of Dr. Paul – December 1, 2016
Greetings everyone, it is with great sorrow that I have to inform you that my father passed away suddenly on November 29th. He was the best dad anyone could ask for. He was always supportive, knew what to say, and made a one-of-a-kind educator. There are no words that could possibly describe how much he’ll be missed by my family, his friends, his colleagues, and me. The phone calls, emails, texts, and videos have been overwhelming. It’s been especially touching for my mom.
This site is part of his legacy. We built it as a team… his knowledge and experience… with the science I had learned through physics. This site and its resources are going nowhere. They will remain as a bastion for the ideals and methods for which he stood.
It’s like a light has gone out. It’s like a light has dispersed, and we’re left trying to fill a void that cannot be filled. Though, we can do what he would want of us. Shine brighter. Shine brighter.
~Adam
.ASHEX14X – March 3, 2014
Hello again, Microbiology Folks,
Adam here with another update. This is long overdue! Today we rolled out ASHEX Version 14X. The new table is available for download in the Non-Fermenter tab. Also, the ASHEX WIP has already been updated to point at the 14X matrix. As of right now, I have not replaced the 12X image logo, but rest assured that the program is using the updated table.
This update means we have expanded to 93 NFB species, represented by 887 clinical isolates.
.Code Rewrite – February 27, 2014
Hello everyone,
I decided it was time to clean up the code a bit. I deployed the new software a few minutes ago, so I figured I would release a statement. There was nothing inherently wrong with the code as it was. Functionally, you should not see any change at all. However, with a little optimization work, I decreased the size of the calculator by another 20%. You may also notice that I have added a search function. This will automatically send the top candidate species to a Google search, allowing for quick access to information.
With any new implementation, however, there is a chance for troubleshooting. Here are a couple potential issues.
1) No matter what test information I put in, the result is P. fulva with a probability of 1.136%.
No fear, this just means your browser is still looking for the old code. Clear your browsing cache/history, and the problem should go away. This can be done from the Tools menu in IE, the History menu in Linux based browsers, and from the advanced settings in Chrome. If you have any issues with this, feel free to shoot me an email.
2) The search function is not working.
This is likely do to the fact that the code wants to open a new tab, which could be disabled by popup blockers / similar prohibitions. Let me know if you are having this kind of error, so I know the extent of the problem.
–Update 2: It seems like this problem is specific to users running Internet Explorer 8.0 browsers. I’ll be looking into a fix.
Thanks! I hope the roll out is a smooth one.
~Adam
.The Cloud MAGI – November 29, 2012
Having trouble pinning an unknown to a point on the GNNFB-Enterics-Vibrio spectrum? Allow me to introduce you to the SAVE MAGI.
This program uses a trio of 18-test matrices to point users to one of the individual WIPs for the likeliest final identification. Just enter the results of the 18 tests and Melchior, Balthazar, and Caspar will put their heads together to compute what they feel is the best option. Click the SAVE Magi option in the AID Cloud menu bar to try it out.
From the office of A.P. Schreckenberger
.ASHEX Web ID Program – July 10, 2012
For those who have trouble downloading programs or have systems incompatible with PIBWin, I have designed a web-based application that will compute ID probabilities without the need to download anything. Use the sidebar menu to access the new ASHEX WIP! The program will list a taxon probability as well as its modal socre. If the modal score falls below the 1.0 threshold, the WIP will display the anomalous results of the top candidate specimen.
Paraphrased from my dad’s email to CLINMICRONET:
Dear Colleagues,
I want to inform you of a new tool for identification of glucose non-fermenting gram-negative bacilli that is available as freeware at www.pschreck.com. This is a computer assisted probability program created by my son Adam, that is linked to the non-fermenter database that I have developed with isolates collected over the past 20 years. The database consists of human clinical isolates and includes rare and unusual isolates from the CDC non-fermenter collection kindly furnished by Maryam Daneshvar and Dannie Hollis. The database includes 45 phenotypic characteristics to 815 individual isolates representing 88 separate species and groups of NFBs. The program allows you to enter as many phenotypic characteristics that are available and calculates likelihood probability, modal score, and provides phenotypic tests that will separate top three taxa when likelihood score is less than 95%. To access, click the WIP tab in the tool bar and enter your reactions on the worksheet. The WIP is a web application that works on any PC, Apple, or smartphone system with an active internet connection. It requires no downloads and will run within your internet browser.
If you would like to access the entire database with all phenotypic reactions click Non-Fermenter ID Tab in the tool bar, then click ASHEX 12X EXCEL File in Step 3. Click read only to open the excel file. If you have any difficulties, you can contact Adam.
We hope you all enjoy this new tool. Thus far, the responses seem to be overwhelmingly positive.
JULY 24 UPDATE:
Dear Microbiology Friends,
In the past week, pschreck.com has introduced the ASHEX WIP – an online tool that helps identify gram-negative, non-fermenting bacilli (GNNFB). Over the course of the last few days, with the help of Brent Barrett, Enterobacteria and Vibrio species WIPs have been developed. Together, with the ASHEX WIP, these programs mark the start of the AID Cloud. The AID Cloud is a set of online identification applications meant to help end-users (such as laboratory specialists) ID clinical isolates.
In the sidebar menu, there are a few WIPs that comprise this set of utilities. The 'Which WIP? AID Cloud' link will take you to a better description of the AID Cloud.
These matrices do not communicate with one another, so users do have to have a general idea as to which WIP they want to use. In other words, the Enterics WIP will not contain information from the ASHEX WIP and vice versa. Feel free to contact Adam with any questions. I hope you all find use for these tools in your labs.
~A.P. Schreckenberger
.Welcome to PSCHRECK.com – May 26, 2011
Welcome to Dr. Paul Schreckenberger’s Clinical Microbiology webpage and the home of the ASHEX Glucose Non-Fermenting Bacteria ID Database. This site was founded in 2004 to provide a home for our project, the ASHEX matrix, which identifies glucose non-fermenting bacteria with a set array of phenotypic tests.

Enjoy your stay,
A.P. Schreckenberger
